GATK Base Recalibration on Bam¶
GATKBaseRecalBQSRWorkflow · 1 contributor · 2 versions
No documentation was provided: contribute one
Quickstart¶
from janis_bioinformatics.tools.common.gatkbasecalbam import GATKBaseRecalBQSRWorkflow_4_1_3 wf = WorkflowBuilder("myworkflow") wf.step( "gatkbaserecalbqsrworkflow_step", GATKBaseRecalBQSRWorkflow_4_1_3( bam=None, reference=None, snps_dbsnp=None, snps_1000gp=None, known_indels=None, mills_indels=None, ) ) wf.output("out", source=gatkbaserecalbqsrworkflow_step.out)
OR
- Install Janis
- Ensure Janis is configured to work with Docker or Singularity.
- Ensure all reference files are available:
Note
More information about these inputs are available below.
- Generate user input files for GATKBaseRecalBQSRWorkflow:
# user inputs
janis inputs GATKBaseRecalBQSRWorkflow > inputs.yaml
inputs.yaml
bam: bam.bam
known_indels: known_indels.vcf.gz
mills_indels: mills_indels.vcf.gz
reference: reference.fasta
snps_1000gp: snps_1000gp.vcf.gz
snps_dbsnp: snps_dbsnp.vcf.gz
- Run GATKBaseRecalBQSRWorkflow with:
janis run [...run options] \
--inputs inputs.yaml \
GATKBaseRecalBQSRWorkflow
Information¶
URL: No URL to the documentation was provided
| ID: | GATKBaseRecalBQSRWorkflow |
|---|---|
| URL: | No URL to the documentation was provided |
| Versions: | 4.1.2, 4.1.3 |
| Authors: | Jiaan Yu |
| Citations: | |
| Created: | 2020-06-12 |
| Updated: | 2020-06-12 |
Outputs¶
| name | type | documentation |
|---|---|---|
| out | IndexedBam |
Workflow¶
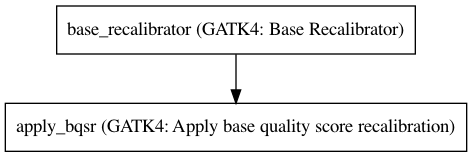
Embedded Tools¶
| GATK4: Base Recalibrator | Gatk4BaseRecalibrator/4.1.3.0 |
| GATK4: Apply base quality score recalibration | Gatk4ApplyBQSR/4.1.3.0 |
Additional configuration (inputs)¶
| name | type | documentation |
|---|---|---|
| bam | IndexedBam | |
| reference | FastaWithIndexes | |
| snps_dbsnp | Gzipped<VCF> | |
| snps_1000gp | Gzipped<VCF> | |
| known_indels | Gzipped<VCF> | |
| mills_indels | Gzipped<VCF> | |
| intervals | Optional<bed> | This optional interval supports processing by regions. If this input resolves to null, then GATK will process the whole genome per each tool’s spec |
Workflow Description Language¶
version development
import "tools/Gatk4BaseRecalibrator_4_1_3_0.wdl" as G
import "tools/Gatk4ApplyBQSR_4_1_3_0.wdl" as G2
workflow GATKBaseRecalBQSRWorkflow {
input {
File bam
File bam_bai
File? intervals
File reference
File reference_fai
File reference_amb
File reference_ann
File reference_bwt
File reference_pac
File reference_sa
File reference_dict
File snps_dbsnp
File snps_dbsnp_tbi
File snps_1000gp
File snps_1000gp_tbi
File known_indels
File known_indels_tbi
File mills_indels
File mills_indels_tbi
}
call G.Gatk4BaseRecalibrator as base_recalibrator {
input:
bam=bam,
bam_bai=bam_bai,
knownSites=[snps_dbsnp, snps_1000gp, known_indels, mills_indels],
knownSites_tbi=[snps_dbsnp_tbi, snps_1000gp_tbi, known_indels_tbi, mills_indels_tbi],
reference=reference,
reference_fai=reference_fai,
reference_amb=reference_amb,
reference_ann=reference_ann,
reference_bwt=reference_bwt,
reference_pac=reference_pac,
reference_sa=reference_sa,
reference_dict=reference_dict,
intervals=intervals
}
call G2.Gatk4ApplyBQSR as apply_bqsr {
input:
bam=bam,
bam_bai=bam_bai,
reference=reference,
reference_fai=reference_fai,
reference_amb=reference_amb,
reference_ann=reference_ann,
reference_bwt=reference_bwt,
reference_pac=reference_pac,
reference_sa=reference_sa,
reference_dict=reference_dict,
recalFile=base_recalibrator.out,
intervals=intervals
}
output {
File out = apply_bqsr.out
File out_bai = apply_bqsr.out_bai
}
}
Common Workflow Language¶
#!/usr/bin/env cwl-runner
class: Workflow
cwlVersion: v1.2
label: GATK Base Recalibration on Bam
doc: ''
requirements:
- class: InlineJavascriptRequirement
- class: StepInputExpressionRequirement
- class: MultipleInputFeatureRequirement
inputs:
- id: bam
type: File
secondaryFiles:
- pattern: .bai
- id: intervals
doc: |-
This optional interval supports processing by regions. If this input resolves to null, then GATK will process the whole genome per each tool's spec
type:
- File
- 'null'
- id: reference
type: File
secondaryFiles:
- pattern: .fai
- pattern: .amb
- pattern: .ann
- pattern: .bwt
- pattern: .pac
- pattern: .sa
- pattern: ^.dict
- id: snps_dbsnp
type: File
secondaryFiles:
- pattern: .tbi
- id: snps_1000gp
type: File
secondaryFiles:
- pattern: .tbi
- id: known_indels
type: File
secondaryFiles:
- pattern: .tbi
- id: mills_indels
type: File
secondaryFiles:
- pattern: .tbi
outputs:
- id: out
type: File
secondaryFiles:
- pattern: .bai
outputSource: apply_bqsr/out
steps:
- id: base_recalibrator
label: 'GATK4: Base Recalibrator'
in:
- id: bam
source: bam
- id: knownSites
source:
- snps_dbsnp
- snps_1000gp
- known_indels
- mills_indels
- id: reference
source: reference
- id: intervals
source: intervals
run: tools/Gatk4BaseRecalibrator_4_1_3_0.cwl
out:
- id: out
- id: apply_bqsr
label: 'GATK4: Apply base quality score recalibration'
in:
- id: bam
source: bam
- id: reference
source: reference
- id: recalFile
source: base_recalibrator/out
- id: intervals
source: intervals
run: tools/Gatk4ApplyBQSR_4_1_3_0.cwl
out:
- id: out
id: GATKBaseRecalBQSRWorkflow